Capturing and connecting research objects: A pitch for @sciencehackday
Jon Eisen asked a question on Friendfeed last week that sparked a really interesting discussion of what an electronic research record should look like. The conversation is worth a look as it illustrates different perspectives and views on what is important. In particular I thought Jon’s statement of what he wanted was very interesting:
I want a system where people record EVERYTHING they are doing in their research with links to all data, analyses, output, etc [CN – my italics]. And I want access to it from anywhere. And I want to be able to search it intelligently. Dropbox won’t cut it.
This is interesting to me because it maps onto my own desires. Simple systems that make it very easy to capture digital research objects as they are created and easy-to-use tools that make it straightforward to connect these objects up. This is in many ways the complement of the Reseach Communication as Aggregation idea that I described previously. By collecting all the pieces and connecting them up correctly we create a Research Record as Aggregation, making it easy to wrap pieces of this up and connect them to communications. It also provides a route towards bridging the divide between research objects that are born digital and those that are physical objects that need to be represented by digital records.
Ok. So so much handwaving – what about building something? What about building something this weekend at ScienceHackDay? My idea is that we can use three pieces that have recently come together to build a demonstrator of how such a system might work. Firstly the DropBox API is now available (and I have a developer key). DropBox is a great tool that delivers on the promise of doing one thing well. It sits on your computer and synchronises directories with the cloud and any other device you put it on. Just works. This makes it a very useful entry point for the capture of digital research objects. So Step One:
Build a web service on the DropBox API that enables users (or instruments) to subscribe and captures new digital objects, creating an exposed feed of resources.
This will enable us to captures and surface research objects with users simply dropping files into directories on local computers. Using DropBox means these can be natively synchronised across multiple user computers which is nice. But then we need to connect these objects up, ideally in an automatic way. To do this we need a robust and general way of describing relationships between them. As part of the OREChem project, a collaboration between Cambridge, Southampton, Indiana, Penn State and Cornell Universities and PubChem, supported by Microsoft, Mark Borkum has developed an ontology that describes experiments (unfortunately there is nothing available on the web as yet – but I am promised there will be soon!). Nothing so new there, been done before. What is new here is that the OREChem vocabulary describes both plans and instances of carrying out those plans. It is very simple, essentially describing each part of a process as a “stage†which takes in inputs and emits outputs. The detailed description of these inputs and outputs is left for other vocabularies. The plan and the record can have a one to one correspondence but don’t need to. It is possible to ask whether a record satisfies a plan and alternately given evidence that a plan has been carried out that all the required inputs must have existed at some point.
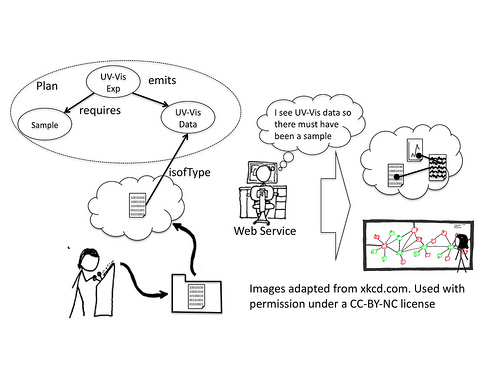
Why does this matter? It matters because for a particular experiment we can describe a plan. For instance a UV-Vis spectrophotometer measurement requires a sample, a specific instrument, and emits a digital file, usually in a specific format. If our webservice above knows that a particular DropBox account is associated with a UV-Vis instrument and it sees a new file of the right type it knows that the plan of a UV-Vis measurement must have been carried out. It also knows which instrument was used (based on the DropBox account) and might know who it was who did the measurement (based on the specific folder the file appeared in). The web service is therefore able to infer that there must exist (or have existed) a sample. Knowing this it can attempt to discover a record of this sample from known resources, the public web, or even by emailing the user, asking them for it, and then creating a record for them.
A quick and dirty way of building a data model and linking it to objects on the web is to use Freebase and the Freebase API. This also has the advantage that we can leverage Freebase Gridworks to add records from spreadsheets (e.g. sample lists) into the same data model. So Step Two:
Implement OREChem experiment ontology in Freebase. Describe a small set of plans as examples of particular experimental procedures.
And then Step Three:
Expand the web service built in Step One to annotate digital research objects captured in Freebase and connect them to plans. Attempt to build in automatic discovery of inferred resources from known and unknown resources, and a system to failover to ask the user directly.
Freebase and DropBox may not be the best way to do this but both provide a documented API that could enable something to be lashed up quickly. I’m equally happy to be told that SugarSync, Open Calais, or Talis Connected Commons might be better ways to do this, especially if someone will be at ScienceHackDay with expertise in this. Demonstrating something like this could be extremely valuable as it would actually leverage semantic web technology to do something useful for researchers, linking their data into a wider web, while not actually bothering them with the details of angle brackets
Related articles by Zemanta
- Dropbox API Lets You Add Cloud Storage to Your Apps (webmonkey.com)
- Why the web of data needs to be social (cameronneylon.net)
- Preview: Freebase Gridworks (simonwillison.net)
- Update: CML, Chem4Word, CheTA, OREChem, Lensfield (wwmm.ch.cam.ac.uk)












Hi Dan, I think Pachube is great for sensors that are running more or less continuously and that kind of thing is definitely part of the ecosystem I see in the longer term. I’m less sure it’s well suited to instruments that are occassionally generating an Excel file, but it would certainly be worth a look.
Could you use Pachube? http://www.pachube.com/
The problem isn't mainly how to share data. The basic problem is that there's few to no incentives for sharing your data in the first place, and many strong incentives against.
License
I am also found at...
Tags
Recent posts
Recent Posts
Most Commented