Frank Gibson of peanutbutter has left a long comment on my post about data models for lab notebooks which I wanted to respond to in detail. We have also had some email exchanges. This is essentially an incarnation of the heavyweight vs lightweight debate when it comes to tools and systems for description of experiments. I think this is a very important issue and that it is also subject to some misunderstandings about what we and others are trying to do. In particular I think we need to draw a distinction between recording what we are doing in the lab and reporting what we have done after the fact. Continue reading “The heavyweights roll in…distinguishing recording the experiment from reporting it”
Semantics in the real world? Part I – Why the triple needs to be a quint (or a sext, or…)
I’ve been mulling over this for a while, and seeing as I am home sick (can’t you tell from the rush of posts?) I’m going to give it a go. This definitely comes with a health warning as it goes way beyond what I know much about at any technical level. This is therefore handwaving of the highest order. But I haven’t come across anyone else floating the same ideas so I will have a shot at explaning my thoughts.
The Semantic Web, RDF, and XML are all the product of computer scientists thinking about computers and information. You can tell this because they deal with straightforward declarations that are absolute. X has property Y. Putting aside all the issues with the availability of tools and applications, the fact that triple stores don’t scale well, regardless of all the technical problems a central issue with applying these types of strategy to the real world is that absolutes don’t exist. I may assert that X has property Y, but what hppens when I change my mind, or when I realise I made a mistake, or when I find out that the underlying data wasn’t taken properly. How do we get this to work in the real world? Continue reading “Semantics in the real world? Part I – Why the triple needs to be a quint (or a sext, or…)”
Proposing a data model for Open Notebooks
‘No data model survives contact with reality’ – Me, Cosener’s House Workshop 29 February 2008
This flippant comment was in response to (I think) Paolo Missier asking me ‘what the data model is’ for our experiments. We were talking about how we might automate various parts of the blog system but the point I was making was that we can’t have a data model with any degree of specificity because we very quickly find the situation where they don’t fit. However, having spent some time thinking about machine readability and the possibility of converting a set of LaBLog posts to RDF, as well as the issues raised by the problems we have with tables, I think we do need some sort of data model. These are my initial thoughts on what that might look like. Continue reading “Proposing a data model for Open Notebooks”
Give me the feed tools and I can rule the world!
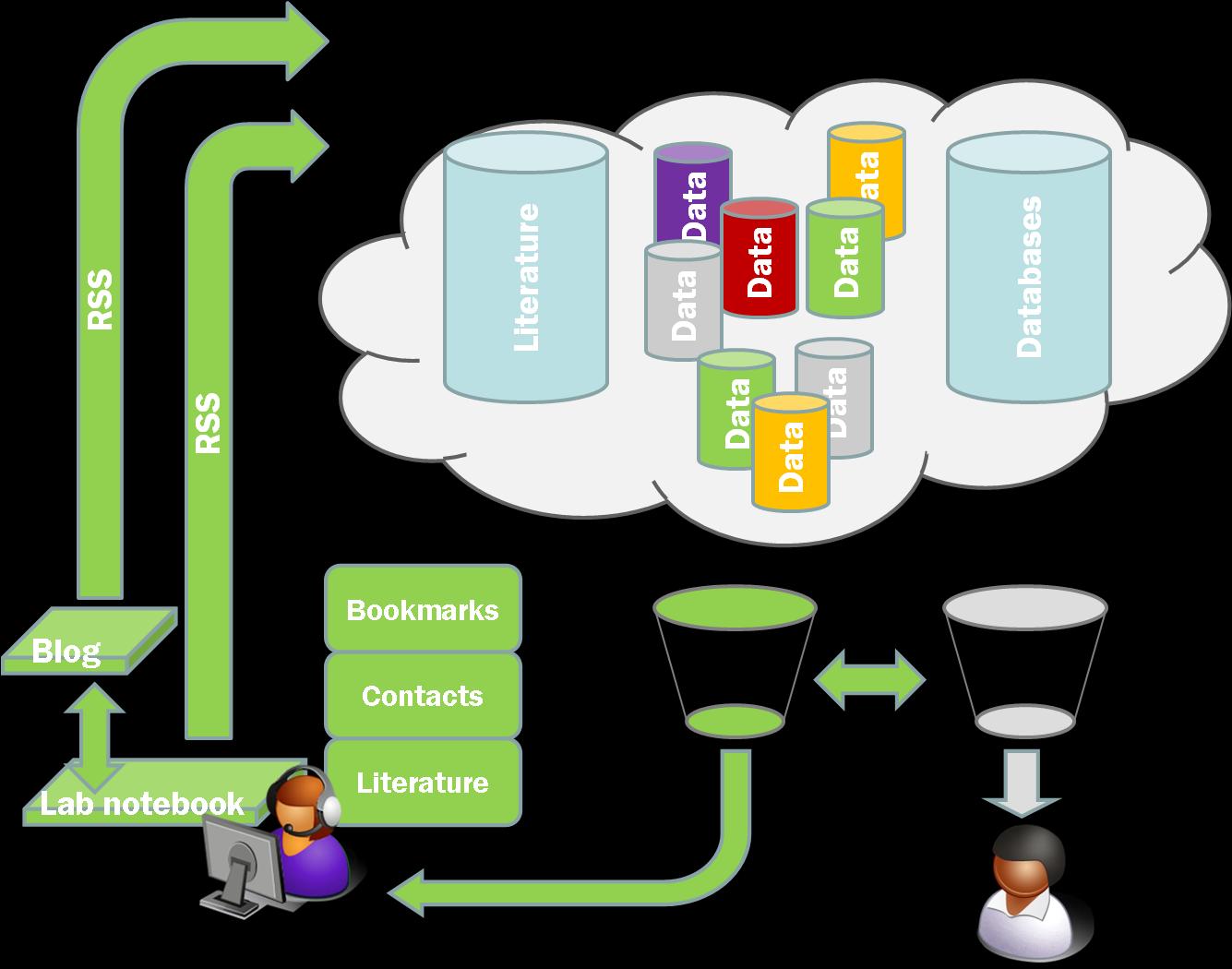
Two things last week gave me more cause to think a bit harder about the RSS feeds from our LaBLog and how we can use them. First, when I gave my talk at UKOLN I made a throwaway comment about search and aggregation. I was arguing that the real benefits of open practice would come when we can use other people’s filters and aggregation tools to easily access the science that we ought to be seeing. Google searching for a specific thing isn’t enough. We need to have an aggregated feed of the science we want or need to see delivered automatically. i.e. we need systems to know what to look for even before the humans know it exists. I suggested the following as an initial target;
‘If I can automatically identify all the compounds recently made in Jean-Claude’s group and then see if anyone has used those compounds [or similar compounds] in inhibitor screens for drug targets then we will be on our way towards managing the information’
The idea here would be to take a ‘Molecules’ feed (such as the molecules Blog at UsefulChem or molecules at Chemical Blogspace) extract the chemical identifiers (InChi, Smiles, CML or whatever) and then use these to search feeds from those people exposing experimental results from drug screening. You might think some sort of combination of Yahoo! Pipes and Google Search ought to do it.
So I thought I’d give this a go. And I fell at the first hurdle. I could grab the feed from the UsefulChem molecules Blog but what I actually did was set up a test post in the Chemtools Sandpit Blog. Here I put the InChi of one of the compounds from UsefulChem that was recently tested as a falcipain 2 inhibitor. The InChi went in as both clear text and as the microformat approach suggested by Egon Willighagen. Pipes was perfectly capable of pulling the feed down, and reducing it to only the posts that contained InChi’s but I couldn’t for the life of me figure out how to extract the InChi itself. Pipes doesn’t seem to see microformats. Another problem is that there is no obvious way of converting a Google Search (or Google Custom Search) to an RSS feed.
Now there may well be ways to do this, or perhaps other tools to do it better but they aren’t immediately obvious to me. Would the availability of such tools help us to take the Open Research agenda forwards? Yes, definitely. I am not sure exactly how much or how fast but without easy to use tools, that are well presented, and easily available, the case for making the information available is harder to make. What’s the point of having it on the cloud if you can’t customise your aggregation of it? To me this is the killer app; being able to identify, triage, and collate data as it happens with easily useable and automated tools. I want to see the stuff I need to see in feed reader before I know it exists. Its not that far away but we ain’t there yet.
The other thing this brought home to me was the importance of feeds and in particular of rich feeds. One of the problems with Wikis is that they don’t in general provide an aggregated or user configurable feed of the site in general or a name space such as a single lab book. They also don’t readily provide a means of tagging or adding metadata. Neither Wikis nor Blogs provide immediately accessible tools that provide the ability to configure multiple RSS feeds, at least not in the world of freely hosted systems. The Chemtools blogs each put out an RSS feed but it doesn’t currently include all the metadata. The more I think about this the more crucial I think it is.
To see why I will use another example. One of the features that people liked about our Blog based framework at the workshop last week was the idea that they got a catalogue of various different items (chemicals, oligonucleotides, compound types) for free once the information was in the system and properly tagged. Now this is true but you don’t get the full benefits of a database for searching, organisation, presentation etc. We have been using DabbleDB to handle a database of lab materials and one of our future goals has been to automatically update the database. What I hadn’t realised before last week was the potential to use user configured RSS feeds to set up multiple databases within DabbleDB to provide more sophisticated laboratory stocks database.
DabbleDB can be set up to read RSS or other XML or JSON feeds to update as was pointed out to me by Lucy Powers at the workshop. To update a database all we need is a properly configured RSS feed. As long as our templates are stable the rest of the process is reasonably straightforward and we can generate databases of materials of all sorts along with expiry dates, lot numbers, ID numbers, safety data etc etc. The key to this is rich feeds that carry as much information as possible, and in particular as much of the information we have chosen to structure as possible. We don’t even need the feeds to be user configurable within the system itself as we can use Pipes to easily configure custom feeds.
We, or rather a noob like me, can do an awful lot with some of the tools already available and a bit of judicious pointing and clicking. When these systems are just a little bit better at extracting information (and when we get just a little bit better at putting information in, by making it part of the process) we are going to be doing lots of very exciting things. I am trying to keep my diary clear for the next couple of months…