Improving on “Access to Research”
Access to Research is an initiative from a 20th Century industry attempting to stave off progress towards the 21st Century by applying a 19th Century infrastructure. Depending on how generous you are feeling it can either be described as a misguided waste of effort or as a cynical attempt to divert the community from tackling the real issues of implementing full Open Access. As is obvious I’m not a neutral observer here so I recommend reading the description at the website. Indeed I would also recommend anyone who is interested to take a look at the service itself.
Building a map of sites
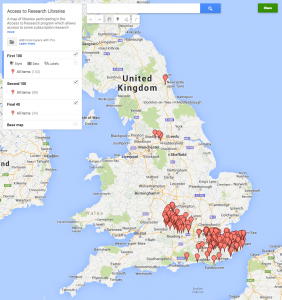
I was interested in possibly doing this myself. In many ways as a sometime researcher who no longer has access to a research library I’m exactly the target audience. Unfortunately the Access to Research website isn’t really very helpful. The Bath public library where I live isn’t a site, nor is Bristol. So which site is closest? Aylesbury perhaps? Or perhaps somewhere near to the places I visit in London. Unfortunately there is no map provided to help find your closest site. For an initiative that is supposed to be focused on user needs this might have been a fairly obvious thing to provide. But no problem, it is easy enough to create one myself, so here it is (click through for a link to the live map).
What I have done is to write some Python code that screen scrapes the Access to Research website to obtain the list of participating libraries, and their URLs. Then my little robot visits each of those library websites and looks for something that matches a UK post code. I’ve then uploaded that to Google Maps to create the map itself. You can also see a version of the code via the IPython Notebook Viewer. Of course the data and code is also available. All of this could easily be improved upon. Surrey County Council don’t actually provide postcodes or even addresses for their libraries on their web pages [ed: Actually Richard Smith and Gary offer sources for this data in the comments – another benefit of an open approach]. I’m sure someone could either fix the data or improve the code to create better data. It would also be nice to use an open source map visualisation rather than Google Maps to enable further re-use but I didn’t want to spend too long on this.
The irony
You might well ask why I would spend a Saturday afternoon making it easier to use an initiative which I feel is a cynical political ploy. The answer is to prove a point. The knowledge and skills I used to create this map are not rare – nor is the desire to contribute to making resources better and more useful for others. But in gathering this data and generating the map I’ve violated pretty much every restriction which traditional publishers want to apply to anyone using “their” work.
What I have done here is Text Mining. Something these publishers claim to support, but only under their conditions and licenses. Conditions that make it effectively impossible to do anything useful However I’ve done this without permission, without registration, and without getting a specific license to do so. All of this would be impossible if this were research that I had accessed through the scheme or if I had agreed to the conditions that legacy publishers would like to lay down for us to carry out Content Mining.
Lets take a look at the restrictions you agree to to use the Access to Research service.
I can only use accessed information for non-commercial research and private study.
Well I work for a non-profit, but this is the weekend. Is that private? Not sure, but there are no ads on this website at least. On the other hand I’ve used a Google service. Does that make it commercial? Google are arguably benefiting from me adding data to their services, and the free service I used is a taster of the more powerful version they charge for.
I will only gain access through the password protected secure service
…oh well, not really, although I guess you might argue that there wasn’t an access restricted system in this case. But is access via a robot bypassing the ‘approved’ route?
I will not build any repository or other archive
…well there would hardly be any point if I hadn’t.
I will not download
…well that was the polite thing to do, grab a copy and process it to create the dataset. Otherwise I’d have to keep hitting the website over and over again. And that would be rude.
I will not forward, distribute, sell
…well I’m not selling it at least…
I will not adapt, modify
…ooops.
I will not make more than one copy…and I will not remove any copyright notices
…well there’s one copy on my machine, one on github, one in however many forks of the repo there are, one on Google…oh and there weren’t any copyright notices on the website. This probably makes it All Rights Reserved with an implied license to view and process the web page but the publisher argument that says I need a license for text mining would mean that I’m in violation of the implied license I would guess. I would argue that as I have only made as many copies required to process and that I have extracted facts to which copyright doesn’t apply I’m fine…but I’m not a lawyer, and this is not legal advice.
I will not modify…any digital rights information.Â
Well there’s one that I didn’t violate! Thank heavens for that. But only because there wasn’t any statement of usage conditions for the site data.
I will not allow the making of any derivative works
…oh dear…
I will not copy otherwise retain [sic] any [copies] onto my personal systems
…sigh
I agree not to rely on the publications as a substitute for specific medical, professional or expert advice.Â
What is ‘expert advice’ I wonder? In any case, don’t rely on the map if you’re in a mad rush.
I could easily go on with the conditions required to sign up for the Elsevier Text Mining program. Again I would either be in clear violation of most of them or it would be difficult to tell. Peter Murray-Rust has a more research-oriented dissection of the problems with the Elsevier conditions in several posts at his blog. Its also quite difficult to tell because Elsevier don’t make those conditions publicly available. The only version I have seen is on Peter’s Blog.
Conclusions
You may feel that I’m making an unfair comparison, that the research content that publishers want to control the use of is different and that the analysis I have done, and the value I have added is different to that involved in using, reading and analysing research. That is both incorrect, and missing the point. The web has brought us a rich set of tools and made it easy for those skilled with them to connect with interesting problems that they can be applied to. The absolutely core question for effective 21st Century research communication is how to enable those tools, skillsets, and human energy to be applied to the outputs of research.
I did this on a rainy Saturday afternoon because I could, because it helped me learn a few things, and because it was fun. I’m one of tens or hundreds of thousands who could have done this, who might apply those skills to cleaning up the geocoding of species in research articles, or extracting chemical names, or phylogenetic trees, or finding new ways to understand the networks of influence in the research literature. I’m not going to ask for permission, I’m not going to go out of my way to get access, and I’m not going to build something I’m not allowed to share. A few dedicated individuals will tackle the permissions issues and the politics. The rest will just move on to the next interesting, and more accessible, puzzle.
Traditional publishers actions, whether this access initiative, CHORUS, or their grudging approach to Open Access implementation, consistently focus on retaining absolute control over any potential use of content that might hypothetically be a future revenue source. This means each new means of access, each new form of use, needs to be regulated, controlled and licensed. This is perfectly understandable. It is the logical approach for a business model which is focussed on monetising a monopoly control over pieces of content. It’s just a really bad way of serving the interests of authors in having their work used, enhanced, and integrated into the wider information commons that the rest of the world uses.











[…] Stop what you’re doing and go read Cameron Neylon’s blog. Specifically, read his new post, Improving on “Access to Researchâ€. […]
Hmm, so for example, being based in Glasgow, this would be a 302 mile round trip to Newcastle.
Once I get to the library, if I manage to get access the such material, I can’t download/save anything.
Think I’ll just stick to reading Open Access papers from right here.
I had concerns about Access To Research in 2012 and in light of this post, my concerns were totally justified:- http://steelgraham.wordpress.com/2013/06/16/the-finch-report/
Also see Mike Taylor’s post “Walk-in access? Seriously?” at SV_POW!: http://svpow.com/2013/11/26/walk-in-access-seriously/
Cameron, you’ve got the same postcode, KT1 2DN, for every library in Surrey (probably because their postcodes aren’t on the the first page of the library site). You’ll need to follow the link with text “Address, travel and accessibility details” for each surreycc library, then scan for the postcode.
This at least expands the map to have good coverage of the home counties (which admittedly is not much better).
If I have time in the week I’ll fix your ipynb.
Richard, yes saw that problem but in the time I had I hadn’t been able to find where the addresses were hidden. A pull request is naturally entirely welcome! I probably won’t get to it for a few days.
Richard, yes saw that problem but in the time I had I hadn’t been able to find the page where the addresses were hidden. A pull request is naturally entirely welcome! I probably won’t get to it for a few days.
Things are much better for me than for Graham. Living in Ruardean, in the Forest of Dean, I have a participating library in Burford, Oxfordshire, only 45 miles away. So for me, it’s only a 90-mile, two-hour round-trip, costing no more than £40 at the HRMC rate of 45p per mile. Why, so long as my time is worth nothing, such a trip could save me money even if I only viewed two paywalled papers that would otherwise have cost the standard $37.95! Although I would of course have to memorise the contents of the papers (or, I suppose, copy them out by hand), since saving is forbidden.
I’ve already said all I have to say about walk-in access (a nineteenth-century solution to a twentieth-century problem. In 2013) so I won’t labour the point here.
But I will say thank you, Cameron, for three things. Firstly, just for providing the map itself, which is very useful. Second, for showing so clearly how the inevitable labyrinth of T&Cs so reliably prevents such things. And third — most important — for showing by example how much more useful data is when set free. As I noted on my own blog, you go beyond critiquing what is, and see what could be.
[…] Cameron Neylon has done a better job of it, but the drive to make research outputs available in libraries is a ruse, and should be called “obfuscating access to research”.  It looks to me like someone in Aberdeen would have to travel to Newcastle (over 250 miles) to find a library where they could read a paper. […]
Maybe the data files on this page would help to feed in the postcodes. http://data.gov.uk/dataset/uk-public-library-contacts-14032012 You can filter it by Surrey and then any web address containing http://www.surreycc.gov.uk . Alternatively the url format for the library addresses follow this structure. http://www.surreycc.gov.uk/people-and-community/libraries/libraries-in-surrey/libraries-and-opening-times/ash-library/ash-library-location-and-access
[…] But no problem, it is easy enough to create one myself, so here it is…” Read on here. Source: […]
[…] the case, points out Advocacy Director for open access journal PLOS Cameron Neylon in a recent blogpost, they would implement full open […]
See this related post http://peerreviewwatch.wordpress.com/2014/02/16/access-to-research-whats-in-it-for-the-publishers-interview/
Hi Cameron, this is Joanna from the Access To Research team at PLS.
Firstly, I just wanted to say that the map you’ve created is a great idea to help users find the nearest participating public library! If possible, I would be really interested in talking to you about how we could adopt and develop this approach for the benefit of those who wish to access the research.
With regard national coverage – as the 2-year pilot has just started, most libraries on the list are still from the technical pilot, which is why some users have had difficulty finding a library close to them. That should change soon though, as since the launch on the 3rd February, over 75% of UK local authorities have expressed an interest in joining the scheme and we are currently busy getting them signed up.
We welcome all feedback on the initiative, and our goal is to work together with everyone who takes an interest in the scheme to make improvements as we go forward, and develop the best possible tools for improving Access to Research. Let me know if you’d be interested in discussing this further.
Joanna Waters
[…] Feb: On his personal blog, Science in the Open, Cameron highlights the absurdity of a new initiative by traditional publishers that has been endorsed by the UK Govt. As part of the FINCH negotiations around Open Access, […]
[…] …one of the motivations I had to get writing again was a request from someone at a traditional publisher to write more because it “was so useful to have a moderate voice to point to”. Seems I didn’t do so well at that with that first post back. […]
[…] libraries across the UK. The launch quickly generated a fair amount of publicity, albeit with equal measures of scorn poured upon […]
[…] Cameron Neylon has rustled up a useful map of the public libraries in the UK set to offer free access to 8,000 commercial paywalled academic ejournals… […]
[…] And it is also these jewels of the digital age that subscription publishers are protecting by trying to restrict text mining. They want you to find their version of the article but only to use it under the conditions they […]
[…] concerns and institutional barriers to change. Cameron Neylon, from PLOS, recently discussed how copyright put up some barriers to his own interesting ideas. Academia is not a nimble beast, and because of it, we are stuck with a lot of scholarly practices […]
License
I am also found at...
Tags
Recent posts
Recent Posts
Most Commented